Peculiar inhibition of human mitochondrial aspartyl-trna synthetaseby adenylate analogs


Biochimie 91 (2009) 596–603
Contents lists available at
Peculiar inhibition of human mitochondrial aspartyl-tRNA synthetaseby adenylate analogs
Marie Messmer Se´bastien P. Blais , Christian Balg Robert Cheˆnevert Luc Grenier , Patrick Lagu¨e ,Claude Sauter Marie Sissler Richard Giege´ Jacques Lapointe , Catherine Florentz
a Architecture et Re´activite´ de l'ARN, Universite´ Louis Pasteur, CNRS, IBMC 15 rue Rene´ Descartes, 67084 Strasbourg Cedex, Franceb De´partement de Biochimie et de Microbiologie, Centre de recherche sur la fonction, la structure et l'inge´nierie des prote´ines (CREFSIP), Faculte´ des sciences et de ge´nie, Universite´Laval, Que´bec, Canada G1V 0A6c De´partement de Chimie, Centre de recherche sur la fonction, la structure et l'inge´nierie des prote´ines (CREFSIP), Faculte´ des sciences et de ge´nie, Universite´ Laval, Que´bec,Canada G1V 0A6
Human mitochondrial aminoacyl-tRNA synthetases (mt-aaRSs), the enzymes which esterify tRNAs with
Received 16 December 2008
the cognate specific amino acid, form mainly a different set of proteins than those involved in the
Accepted 18 February 2009
cytosolic translation machinery. Many of the mt-aaRSs are of bacterial-type in regard of sequence and
Available online 28 February 2009
modular structural organization. However, the few enzymes investigated so far do have peculiarbiochemical and enzymological properties such as decreased solubility, decreased specific activity and
enlarged spectra of substrate tRNAs (of same specificity but from various organisms and kingdoms), as
Human mitochondria
compared to bacterial aaRSs. Here the sensitivity of human mitochondrial aspartyl-tRNA synthetase
Aspartyl-tRNA synthetase
(AspRS) to small substrate analogs (non-hydrolysable adenylates) known as inhibitors of Escherichia coli
and Pseudomonas aeruginosa AspRSs is evaluated and compared to the sensitivity of eukaryal cytosolic
Molecular docking
human and bovine AspRSs. L-aspartol-adenylate (aspartol-AMP) is a competitive inhibitor of aspartyla-tion by mitochondrial as well as cytosolic mammalian AspRSs, with Ki values in the micromolar range (4–27 mM for human mt- and mammalian cyt-AspRSs). 50-O-[N-(L-aspartyl)sulfamoyl]adenosine (Asp-AMS)is a 500-fold stronger competitive inhibitor of the mitochondrial enzyme than aspartol-AMP (10 nM) anda 35-fold lower competitor of human and bovine cyt-AspRSs (300 nM). The higher sensitivity of humanmt-AspRS for both inhibitors as compared to either bacterial or mammalian cytosolic enzymes, is notcorrelated with clear-cut structural features in the catalytic site as deduced from docking experiments,but may result from dynamic events. In the scope of new antibacterial strategies directed against aaRSs,possible side effects of such drugs on the mitochondrial human aaRSs should thus be considered.
Ó 2009 Elsevier Masson SAS. All rights reserved.
within the cytosol and imported . They include the sets of ami-noacyl-tRNA synthetases (aaRSs), ribosomal proteins, translation
Two distinct translational machineries coexist in mammalian
factors and tRNA maturation and modification enzymes. AaRSs are
cells. The mitochondrial machinery is still in the process of char-
the enzymes which catalyze specific esterification of their cognate
acterization. While its 22 tRNAs, 11 mRNAs (2 are polycistronic) that
tRNAs by the corresponding amino acids. Most of the genes
code for 13 proteins, and 2 rRNAs are encoded by the mitochondrial
encoding the human aaRSs have been annotated, demonstrating
(mt) genome, all other macromolecules needed for protein
their distribution into two distinct sets Except for GlyRS
synthesis are coded by the nuclear chromosome, synthesized
and LysRS , mt- and cytosolic-aaRSs (cyt-aaRSs) are encoded bydistinct genes.
In agreement with the endosymbiotic hypothesis for the origin
of mitochondria sequence features and modular organizationof many mt-aaRSs are of bacterial-type and thus differ from the
Abbreviations: aaRS, aminoacyl-tRNA synthetase, with aa for the amino acid in
eukaryotic-type corresponding cyt-aaRSs Biochemical and
three-letter abbreviation; mt, mitochondrial; cyt, cytosolic; aspartol-AMP, aspartol-
enzymatic characterization of an initial set of human bacterial-type
mt-aaRSs revealed however unexpected properties making these
* Corresponding author. Tel.: þ33 3 88 41 70 59; fax: þ33 3 88 60 22 18.
E-mail address: (C. Florentz).
enzymes functionally distinct from their bacterial counterparts. As
0300-9084/$ – see front matter Ó 2009 Elsevier Masson SAS. All rights reserved.
doi:10.1016/j.biochi.2009.02.005
M. Messmer et al. / Biochimie 91 (2009) 596–603
an example, human mt-AspRS and mt-TyrRS share about 40%
2. Materials and methods
sequence identity with the corresponding Escherichia coli enzymes(including strongly conserved functional amino acids) and present
2.1. Materials and enzymes
the same modular organization . The crystallographic structureof mt-TyrRS reveals a three-dimensional fold very similar to that of
Total E. coli tRNA was purchased from Roche Diagnostics and total
Bacillus subtilis and E. coli TyrRSs . However, both mt-aaRSs
calf liver tRNA from Novagen. L-[2,3-3H]aspartic acid (specific
aminoacylate their substrates with 10- to 40-fold less efficiency
activity 34 Ci/mmol) was from GE Healthcare. Aspartol-AMP and
than the corresponding E. coli aaRSs . These mt-enzymes require
Asp-AMS were synthesized as reported Ni-NTA resin was from
restricted sets of identity elements within their cognate tRNAs
Qiagen Inc. Human (Homo sapiens) mt-AspRS was previously cloned
compared to bacterial AspRSs or TyrRSs . Finally, both mt-
into pQE70 vector that introduces a poly-His tag to the C-terminus of
AspRS and mt-TyrRS likely have an enlarged spectrum of possible
the expressed protein. Overproduction and purification steps were
tRNA substrates, as first observed after comparing E. coli aaRSs with
conducted as described Human cyt-AspRS was a kind gift of
homologous bovine enzymes . Indeed mitochondrial enzymes
M. Frugier (Stasbourg). Bovine (Bos taurus) cyt-AspRS was purchased
aminoacylate tRNAs of same specificity from a large range of
from Bio S&T Inc. (Montreal, Canada) as a mixture of different ami-
organisms, while most bacterial enzymes recognize and amino-
acylate only their own tRNA.
Aminoacyl-tRNA synthetases have been subjected to signifi-
2.2. Aminoacylation and inhibition assays
cant evolutionary divergence, so that selective inhibition ofbacterial enzymes appears as a valuable strategy for the
Aspartylation assays in the presence of human mt-AspRS were
production of new antibiotics (reviewed in Refs. Such
carried out in 50 mM HEPES-NaOH pH 7.5, 2.5 mM ATP, 12 mM
antibiotics are expected to have strong negative effects on
MgCl2, 25 mM KCl, 0.2 mg/ml BSA, 1 mM spermine and 40 mM total
pathogenic bacteria, but should not affect the human host.
E. coli tRNA. For establishing the Km for aspartate, this substrate was
Pseudomonic acid (mupirocin) is the first known effective anti-
added at concentrations ranging from 0.7 to 40 mM. The reaction
biotic of this type and inhibits IleRSs from Gram positive (e.g.
was initiated by adding the pre-warmed enzyme at 37 �C to a final
Staphylococcus aureus) and Gram negative (e.g. Neisseria menin-
concentration of 62.5 nM. The amount of aspartyl-tRNA formed
gitidis) bacteria with a 8000-fold higher affinity than for
was determined by the radioactivity present in 5% trichloroacetic
mammalian cyt-IleRS . This natural product is a stable
acid precipitates of reaction mixture aliquots, as previously
adenylate analog and is in clinical use Beside IleRS, many
described Initial reaction rates were determined by measuring
other aaRSs are inhibited by adenylate derivatives, of synthetic
[3H]aspartyl-tRNA formed in 5 ml aliquots (from a total volume of
and in a few cases of natural origin, that can be considered as
50 ml) taken at 1 min intervals over 6 min. Inhibition constants (Ki)
potential drugs targeting aaRSs .
were determined at the aspartate concentration corresponding to
We have previously synthesized aspartyl-adenylate analogs
the Km value and the inhibitors aspartol-AMP and Asp-AMS were
and established that they have inhibitory effects on bacterial
added at various concentrations from 0.5 to 100 mM and from 1 to
aaRSs as tested on E. coli and Pseudomonas aeruginosa AspRSs
50 nM, respectively, to reaction media pre-heated to 37 �C, 2 min
Here, the effect of Asp-AMS and aspartol-AMP () is explored on
before addition of the synthetase. The error range was 15% for
human mt-AspRS as well as on mammalian cytosolic AspRSs
(human and bovine). The inhibition produced by Asp-AMS and
Aspartylation assays performed in the presence of human cyt-
aspartol-AMP on the activity of the three enzymes was investigated
AspRS were carried out as for human mt-AspRS but with 80 mM total
and compared with the effect produced on bacterial AspRSs.
calf tRNA and 0.5 nM enzyme. To determine the Km for aspartate for
Functional studies were completed by computer-assisted docking
the human cyt-AspRS, this substrate was added to final concentra-
of the adenylates in the catalytic site of the diverse AspRSs. Data
tions ranking from 2.5 to 120 mM. Aliquots of 5 ml were taken from
reveal differences between the three types of enzymes (bacterial,
a total volume of 50 ml, at 2 min intervals over 12 min and treated as
mt- and cyt-eukaryal) and strikingly highest sensitivity of the
described above. Establishment of Ki for aspartol-AMP and Asp-AMS
mitochondrial enzyme to both inhibitors. They further support
was set to 24 mM of aspartate (Km value) and was done as described
functional differences between the bacterial-type human mt-
above but with inhibitor concentrations varying from 5 mM to
AspRS and bacterial AspRSs. These functional peculiarities are not
100 mM and from 50 nM to 1 mM, respectively. The error range was
due to striking structural idiosyncrasies in the catalytic domain of
10% for triplicate experiments.
the AspRSs, in particular of human mt-AspRS, as suggested bydocking of the adenylate in the catalytic sites. Structure–function
2.3. Determination of inhibition type and constant (Ki)
relationships and the implications for medical research dedicatedto the discovery of new antibiotics using aaRSs as targets will be
The Km values of human cyt-AspRS and human mt-AspRS for the
aspartate substrate were calculated from Michaelis–Menten plots.
Fig. 1. Comparison of the chemical structures of aspartyl-adenylate and its two analogs, L-aspartol-adenylate and 50-O-[N-(L-aspartyl)sulfamoyl]adenosine.
M. Messmer et al. / Biochimie 91 (2009) 596–603
The rate ‘vi' of the aminoacylation reaction in the presence of
Docking of the natural aspartyl-adenylate and of two analogs
inhibitor at various concentrations [I] is given by the following
(Asp-AMS and aspartol-AMP) was performed using AUTODOCK
3.0.5 . Hydrogen atoms were added to proteins and ligandsusing
Gast-Eigen charges were computed for the ligand partial atomic
½S� þ Kmð1 þ ð½I�=KiÞÞ
Three-dimensional grids of interaction energies based on the
When reactions are conducted at the amino acid concentration
macromolecular target using the AMBER force field were calculated
corresponding to the Km value and in the presence of saturating
using AutoGrid. The cubic grid box of 60 Å size (x, y, z) with
concentrations of ATP and tRNA, Eq. can be rearranged as Eq.
a spacing of 0.375 Å and grid maps were centered on the respective
with the ratio vi/v0 (where v0 is the rate in the absence of inhibitor
AspRS active sites. Automated docking studies were carried out to
under the same substrate concentrations) expressed as a function
evaluate the binding free energy of the inhibitors within the
of [I]. This equation illustrates competitive inhibition with one
macromolecules. The GA-LS search algorithm (algorithm with local
binding site for the inhibitor and can be simplified in Eq.
search) was chosen to search for the best conformers. The param-
eters were set using the software ADT. For all docking parameters,
default values were used with 30 independent docking runs for
each docking case. Each three-dimensional model was used as
½S� þ Kmð1 þ ð½I�=KiÞÞ
a rigid scaffold, but the three ligands benefited from a full freedomwith respect to their flexibility to be able to adapt to the catalytic
groove. A theoretical Ki value was derived from the calculated
binding free energy [Ki ¼ exp(DGbinding/RT)]. An average pKi value
Curve-fitting of the data was made with the Sigmaplot software
(pKi ¼ �log Ki) is given for all trials falling within 2 Å rmsd from the
(SPSS Inc) and was used to identify the type of inhibition and to
position of the natural adenylate in crystallographic structure of
E. coli ternary complex. The figure presenting docking results was
i values. Inhibition type of Asp-AMS with respect
to aspartate for both human AspRSs was further confirmed by
prepared with PyMOL (DeLano Scientific LLC, CA).
the determination of the apparent Km for aspartate in thepresence of several concentrations of this inhibitor (from 0 to
40 nM for human mt-AspRS and from 0 to 1.2 mM for human cyt-AspRS) and under fixed and saturating concentrations of the two
3.1. Activity of human and bovine AspRSs inhibited by aspartyl-
other substrates (2.5 mM ATP and 40 mM E. coli total tRNA for
adenylate analogs
human mt-AspRS or 80 mM total calf tRNA for human cyt-AspRS).
As a prerequisite to the search of the inhibition of adenylates on
AspRS activity (see ) we determined the
2.4. Docking of adenylate and analogs
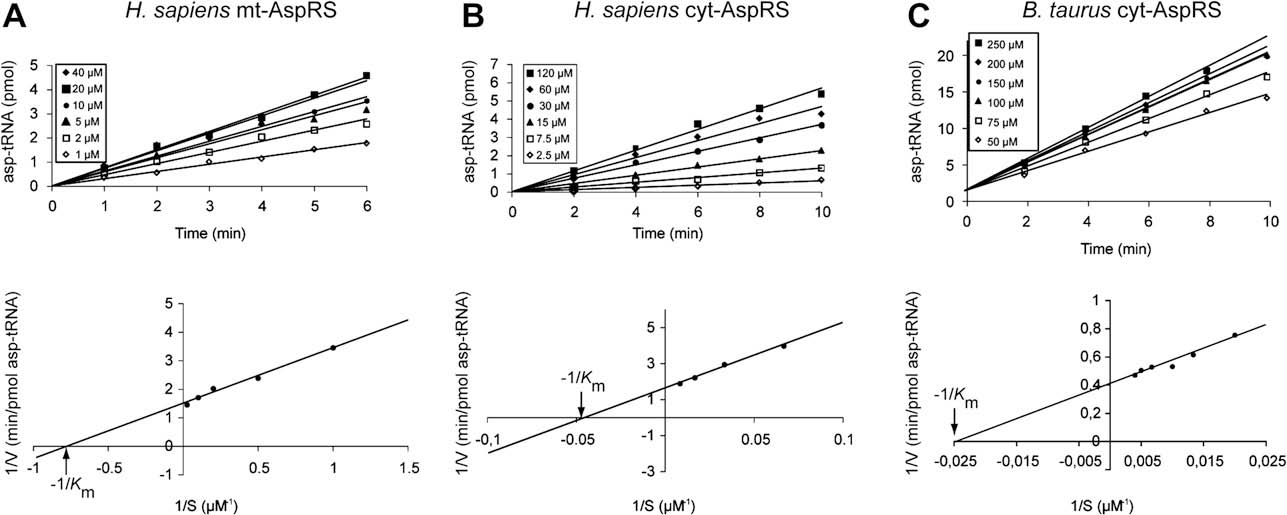
Km for aspartate of human cyt- and mt-AspRSs and for bovine cyt-AspRS. Initial rates of aspartylation, established under saturating
Three-dimensional models of candidate AspRSs were derived
tRNA and ATP concentrations were obtained for aspartate
from crystallographic structures of close relatives in complex with
concentrations ranging from 1 to 40 mM (mt-AspRS) and 2.5 to
tRNA. Yeast binary complex (1ASY.pdb – Ref. ) and the E. coli
120 mM (cyt-AspRSs) , top). Analysis of the Lineweaver–Burk
ternary complex (1CA0.pdb – Ref. were used to model bovine
plots (, bottom) yielded similar Km values for the two cytosolic
and human cyt-AspRSs and bacterial-type enzymes (P. aeruginosa
AspRSs (24 mM and 37 mM for the human and bovine enzymes,
and human mitochondria, respectively). The three-dimensional
respectively) and a strikingly lower value (1.5 mM) for the human
model of the mt-AspRS was built using modeler as described
previously whereas others were generated using the web-
The inhibition constants (Ki) of Asp-AMS and aspartol-AMP have
based SWISS-MODEL workspace .
been determined with respect to aspartate for the three AspRSs at
Fig. 2. Determination of the apparent Km values for aspartate for H. sapiens mt-AspRS (A), H. sapiens cyt-AspRS (B) and for B. taurus cyt-AspRS (C).
M. Messmer et al. / Biochimie 91 (2009) 596–603
fixed and saturating concentrations of ATP and tRNA and at Km
coordinates from E. coli complex used as a template for homology
concentration for the aspartic acid. Kinetic data were displayed as
modeling did contain this ligand. The difference of 3 theoretical pKi
normalized initial rates of tRNA aminoacylation (vi/v0,) as a func-
units between bacterial- and eukaryotic-type systems may also be
tion of inhibitor concentration Ii (with vi being the initial rates in
linked to the resolution of the original template, the E. coli structure
the presence of inhibitor and v0 the rate in the absence of inhibitor).
determined with a higher accuracy (2.3 vs 2.9 Å resolution) giving
If inhibitions are competitive, as can be anticipated for adenylate
the highest docking scores. Beside these technical aspects, the most
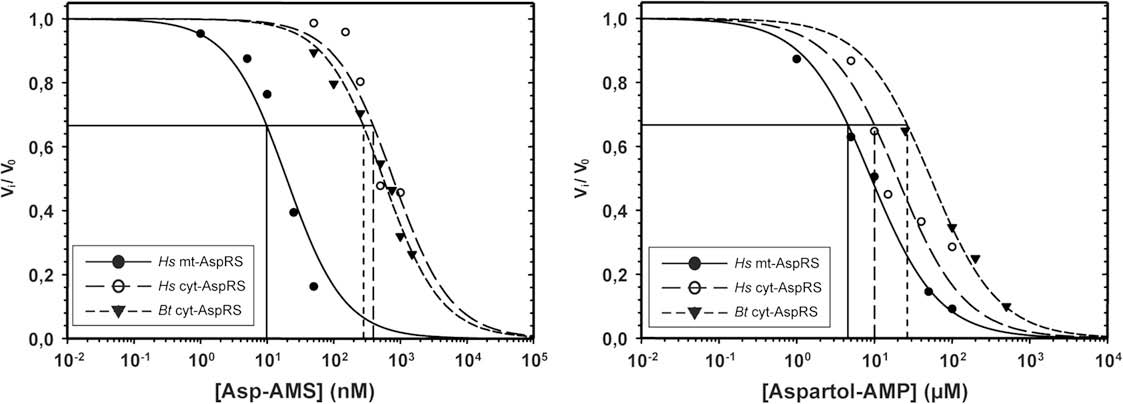
analogs, experimental data should fit on sigmoidal curves (see
striking feature is that pKi values obtained for a given AspRS do not
dramatically vary from one ligand to the others. Moreover, no
computed for aspartol-AMP fit perfectly with the experimental
significant behavior difference is detected between AspRSs
points indicating that the inhibition is indeed competitive
belonging to the same group.
for the three enzymes with Ki values ranging from 4.6 to 27 mM
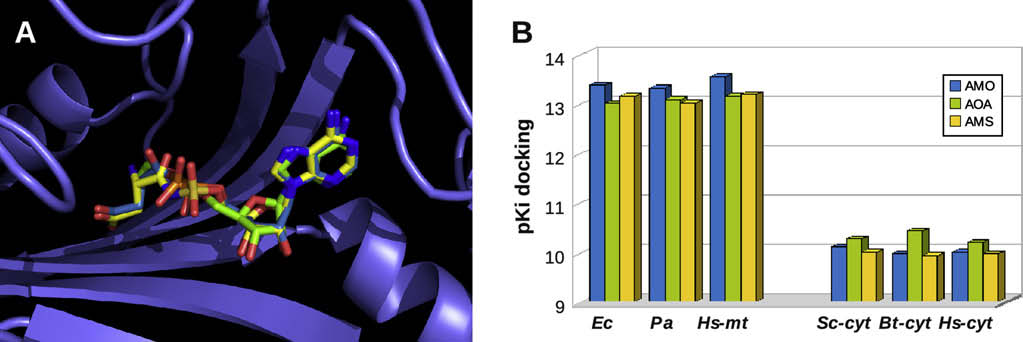
As an illustration, shows the locations of aspartyl-adeny-
). In the experiments done in the presence of Asp-AMS, the
late, aspartol-AMP and Asp-AMS superimposed in the active site
fit is less perfect, especially in the case of the human mt-AspRS.
human mt-AspRS (in the homology model of E. coli AspRS). The
Assuming inhibitions are competitive, extracted Ki values are quite
docking suggests excellent superimpositions of the adenine
similar for the two mammalian cyt-AspRSs (390 and 280 nM for the
(A, right) and aspartate , left) moieties at the distal
human and bovine enzymes) and 9.8 nM for the mt-AspRS. These
extremities of the adenylate molecules and slight changes in the
values are 2–3 orders of magnitude below those measured for
orientation of the ribo-phosphate and ribo-sulfamoyl groups in the
central part of these molecules.
To verify whether the deviations from the theoretical curve with
Asp-AMS originate from experimental errors or result from a more
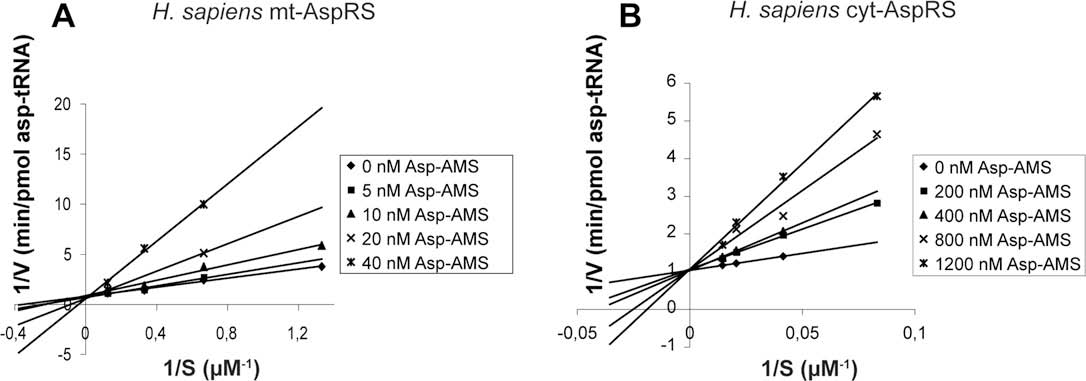
complex kinetic behavior, we undertook a classical Lineweaver–Burk analysis for the human mitochondrial and cytosolic enzymes
4.1. General considerations
Apparent Km values for aspartate have been determinedunder large ranges of inhibitor concentrations. All lines cross the y-
Aminoacyl-tRNA synthetases catalyze the esterification of their
axis into a single point that corresponds to 1/Vmax, conclusively
cognate tRNA with the specific amino acid in a two-step process. In
demonstrating that inhibition by Asp-AMS is of competitive type
the first step, the amino acid is recognized by the enzyme and
with respect to aspartate both for human cyt-AspRS and mt-AspRS.
reacts with ATP to form an enzyme-bound mixed anhydride (aa-
Kinetic parameters in these experiments are very close to those
AMP or aminoacyl-adenylate) with release of pyrophosphate .
In this intermediate, the high-energy anhydride bond activates thecarboxyl group of the amino acid. In the second step, the activated
3.2. Molecular docking of aspartyl adenylate and analogs in the
amino acid is transferred to the 30-terminal adenosine of the cor-
active site of AspRSs
responding tRNA to form aminoacyl-tRNA and AMP (reviewed inRefs. This overall mechanism applies to both class I and
Three-dimensional models of four AspRSs (from P. aeruginosa,
class II aaRSs (reviewed in Refs. ). While the overall func-
human mitochondria, human and bovine cytoplasms) were
tioning of aaRSs is essentially conserved in evolution, one notes
generated to study the binding of aspartyl-adenylate and its
idiosyncrasies when comparing properties of aaRSs from phylo-
analogs (Asp-AMS and aspartol-AMP) by molecular docking. These
genetically distant species or organelles and this opens the
three-dimensional models were based on X-ray structures from
possibility to find or design species-selective inhibitors of aaRSs.
E. coli and yeast AspRS:tRNAAsp complexes (see
Here we focus on human mt-AspRS for a better understanding of
). Individual monomers were considered (i.e. one active
its functional and structural idiosyncrasies, especially in regard of
site) in the absence of tRNA. In order to get comparative scores,
inhibition by small substrate analogs targeting its catalytic site. For
docking trials were performed for each ligand, both on the original
comparative purposes, two novel mammalian cyt-AspRSs (human
X-ray structures and on the four models (with protein backbone
and bovine) were studied for their behavior to interact with aspartic
and side chain orientation maintained as in the reference X-ray
acid and two adenylate analogs. reports the Km and Ki values
structures). The results are presented in In the case of
for the three mammalian enzymes and compares these values with
bacterial-type enzymes, the natural adenylate systematically gives
those previously determined for E. coli and P. aeruginosa AspRSs .
a slightly better score, probably due to a structural bias: the X-ray
Remarkable variations are observed that are best visualized in the
Fig. 3. Inhibition kinetics with Asp-AMS (left) and aspartol-AMP (right) of H. sapiens mt-AspRS, H. sapiens cyt-AspRS and B. taurus cyt-AspRS. Abbreviations used: Hs, Homo sapiens;Bt, Bos taurus.
M. Messmer et al. / Biochimie 91 (2009) 596–603
27 mM) and for Asp-AMS (390 and 280 nM). Over the 5 enzymes
Kinetic parameters Km of aspartyl-adenylate and Ki of non-hydrolysable analogs for
considered, human mt-AspRS is the most sensitive enzyme towards
mammalian and bacterial AspRSs. Abbreviations used: Pa, Pseudomonas aeruginosa;
each inhibitor with a Ki of 4.6 mM for aspartol-AMP and of 9.8 nM
Ec, Escherichia coli; Hs, Homo sapiens; Bt, Bos taurus. n.d. stands for non-determined.
for Asp-AMS ().
Aspartate Km (mM)
Aspartol-AMP Ki (mM)
4.3. H. sapiens mitochondrial AspRS is more sensitive to adenylates
than bacterial AspRSs
The data of and highlight distinct behaviors for the
three families of enzymes considered (bacterial, bacterial-type,
eukaryal). Considering either K
Experimental data taken from Ref.
m values for the natural substrate
aspartate or Ki values of the inhibitors, the mitochondrial enzymebehaves apart from the four other AspRSs discussed here. Not only
histogram comparing the inverse of the Km and Ki values ().
is aspartic acid retained with the best relative affinity for this
Human mt-AspRS presents the most atypical functional behavior
enzyme (assuming that the inverse of Km values is representative of
deviating significantly from what observed with other AspRSs. It is
the affinity) but also the adenylate analogs do present the highest
common sense to believe that the functional differences are due to
relative inhibitory properties.
structural idiosyncrasies of the different AspRSs, but as shown in
Mt-AspRS displays a much higher affinity for aspartic acid than
other tRNA aminoacylation systems, large functional differences
the two cyt-AspRSs (16–25-fold) and than the two bacterial AspRSs
could originate from faint structural effects . For interpreta-
(60–70-fold). This markedly small Km value is in support of
tion of the present data it should be kept in mind that all the above
distinct kinetic properties for the enzyme families considered and
results were obtained by kinetic analyses conducted in the presence
especially of the mitochondrial bacterial-type enzyme as compared to
of tRNA, and that former experiments with yeast AspRS have shown
the two other families. Bacterial synthetases (E. coli and P. aeruginosa)
that tRNAAsp significantly increases the affinity of aspartyl-adenylate
present the poorest affinity for their amino acid substrate, while
for the synthetase .
the two eukaryal enzymes present a 3–4-fold better affinity.
In regard to inhibitors, aspartol-AMP presents a 2–6-fold lower Ki
4.2. Aspartol-AMP and Asp-AMS are competitive inhibitors of
for mt-AspRS than for the human or bovine cyt-AspRSs and about
H. sapiens and B. taurus AspRSs
10-fold lower than the bacterial AspRSs. Asp-AMS has also thehighest inhibitory effect for the mitochondrial enzyme, but it
Aspartol-AMP and Asp-AMS are analogs of aspartyl-adenylate,
remains close to those measured for E. coli AspRS but distinguishes
the natural derivative formed by AspRS in the presence of aspartic
strongly from the cyt-enzymes. In summary, whatever the inhibitor,
acid and ATP, during the first step of the aminoacylation reaction.
it has the strongest effect on the mitochondrial enzyme. This
Aspartol-AMP differs from aspartyl-adenylate in converting an
enzyme distinguishes thus from both other families of considered
aminoacyl-adenylate into an aminoalcohol-adenylate, while Asp-
AspRSs, namely bacterial and eukaryal cytosolic AspRSs. Note that
AMS has a sulfamoyl function In a previous work, it was
both these families present an inverted reactivity, with inhibition by
shown as anticipated, that both molecules are competitive inhibi-
aspartol-AMP more important with the eukaryal enzymes and
tors of aspartate in bacterial AspRSs (E. coli and P. aeruginosa)
inhibition by Asp-AMS more important with the prokaryal enzymes.
. While aspartol-AMP is a weak inhibitor for these twoAspRSs with Ki values in the micromolar (mM) range, Asp-AMS is
4.4. Search for structure–function relationships
a strong inhibitor with Ki in the nanomolar (nM) range ). Inthe present work, that extends the analysis to eukaryal cyt-AspRSs
Two aspects have to be considered here. First, the functional
(human and bovine) and to human mt-AspRS, both adenylate
difference between the human mt-AspRS and the two bacterial
analogs behave also as competitive inhibitors. Inhibition constants
AspRSs from E. coli and P. aeruginosa, given the fact that the human
(Ki) of aspartol-AMP remain in the mM range and those of Asp-AMS
enzyme is of bacterial-type and thus structurally similar to
remain in the nM range, as was the case for the bacterial enzymes
bacterial AspRSs. Second, the strong inhibitory effect produced by
. Interestingly, both cytosolic mammalian AspRSs (human
Asp-AMS with an affinity about three orders of magnitude higher
and bovine) have about the same Ki for aspartol-AMP (10 and
than the closely related aspartol-AMP ). In the absence of
Fig. 4. Inhibition with Asp-AMP of the aminoacylation activity of H. sapiens mt-AspRS (A) and cyt-AspRS (B). Experiments have been performed in the presence of different fixedconcentrations of Asp-AMS.

M. Messmer et al. / Biochimie 91 (2009) 596–603
Fig. 5. Docking of aspartyl-adenylate and of two analogs in the active site of AspRSs. (A) Example of the best docking solution (i.e. highest binding energy) for the natural adenylate(or AMO with carbon atoms in medium blue), aspartol-AMP (or AOA, in green) and Asp-AMS (or AMS, in yellow). The protein backbone (with the antiparallel b-sheet characteristicof class II aaRSs) is represented in blue. Small variations are observed at the connection of the two adenylate moieties, whereas the position of the aspartate side chains and of theadenine ring is almost conserved. (B) Average docking scores obtained for the three substrates (same color code) in the active site of five AspRSs. Scores are indicated in theoreticalpKi values based on computed binding energies (see ). (For interpretation of color in this figure, the reader is referred to the web version of this article.)
crystallographic structures of human mt-AspRS in its apo and
catalysis (see legend to ) and only 3 differ in the two enzymes,
liganded versions, as well as of the other AspRSs investigated in this
namely Ile536, Ile581 and Leu583 in mt-AspRS, replaced respec-
work, except the E. coli enzyme , a structural analysis of AspRSs
tively by Phe533, Val483 and Leu531 in E. coli AspRS. These amino
completed by docking studies of the adenylates in the active site of
acids are not predicted to make energetically favorable bounds
AspRSs can be useful.
with the adenylate, and in addition were not identified by crystal-
AspRSs are modular proteins that belong to class IIb aaRSs. Their
lography to contribute to adenylate binding in E. coli AspRS. This
catalytic core encompasses a seven-stranded antiparallel b-sheet,
suggests that the functional differences between the two AspRSs
surrounded by a-helices that encompass the three class II signature
are due to subtle structural effects and are not accounted by sole
motifs. This fold is common to all class II aaRSs and differs from the
thermodynamic binding features but are also kinetically driven
Rossmann-fold of class I aaRSs The b-sheet offers a platform
with induced fit and indirect effects. Such an interpretation finds
where adenylates are formed. The overall structure and the cata-
support from a mutational analysis of the active site of yeast AspRS
lytic core of AspRSs are roughly conserved in evolution, but present
that identified 23 functionally important amino acids by a genetic
idiosyncrasies specific to phylogenic kingdoms (lower and higher
selection method. Among these amino acids located around the
eukarya, bacteria, archaea and organelles) (reviewed in Ref.
ATP binding site, 10 act indirectly and were not identifiable by
Interestingly, crystallographic structures tell us that the inter-
crystallography .
action of aspartyl-adenylate with AspRSs is essentially the same
In conclusion this analysis suggests that the Km and Ki variations
than with free aspartate and the adenosine moiety of ATP. Indeed,
observed in the test tube are not only the consequence of the
as found with E. coli AspRS (), the AMP moiety of aspartyl-
architecture of the active site itself and of the direct atomic envi-
adenylate is positioned in a class II conserved manner, with
ronment of the ligands, but also rely on the dynamics of ligand
interactions of the a-phosphate with conserved Arg217 (Arg266 in
binding, tRNA and small substrates, and the associated conforma-
mt-AspRS) from motif 2. Further, recognition of the a-carboxyl and
tional changes. In this process, adaptability of the flexible tRNA
a-amino groups of aspartate by conserved AspRS residues is also
molecule on the protein will likely be crucial . Deciphering
class II characteristic, but with a system-specific interaction of the
these subtleties of human mt-AspRS will require more functional
side chain carboxylic group with Lys198, Arg489 (Lys247 and
and structural work. In regard to functional investigation, it should
Arg542 in mt-AspRS) whose basic side chains are stabilized by saltbridges with Asp233, Glu235 (would be Asp282 and Glu284 in mt-AspRS). Comparison between the E. coli AspRS structure containingaspartyl-adenylate and the apo structure shows that there is noconformational change whatsoever of the four conserved residuesLys198, Asp233, Glu235 and Arg489 (correspond to Lys247, Asp282,Glu284, Arg542 in mt-AspRS) from the catalytic domain uponaspartate binding . This implies a ‘‘lock-and-key'' recognition ofthe preformed adenylate analogs that is also found in archaealAspRS from Pyrococcus kodakaraensis and contrasts with theinduced fit occurring upon recognition of ATP with conformationalchanges in the active site of the E. coli enzyme, in particular atArg217.
On the other hand, sequence analysis of human mt-AspRS
together with modeling of its three-dimensional structure (basedon the structure of E. coli AspRS:tRNAAsp:adenylate ternarycomplex, see and docking of the adeny-lates, indicates similar interaction patterns of the adenylates (see). The amino acids in human mt-AspRS and E. coli AspRSidentified by the docking procedure to make energetically favorable
Fig. 6. Histogram comparing the (1/Km) values of aspartate and the (1/Ki) values of thetwo non-hydrolysable aminoalcohol (aspartol-AMP) and sulfamoyl (Asp-AMS) deriv-
bonds with the adenylates or to be in vicinity of the adenylate in the
atives for five different AspRSs. Abbreviations used: Pa, Pseudomonas aeruginosa; Ec,
active cavity of the AspRSs are shown in Among these 24
Escherichia coli; Hs, Homo sapiens; Bt, Bos taurus. Experimental data for Pa and Ec
amino acids, 10 were identified by crystallography to play a role in
AspRSs are taken from Ref.
M. Messmer et al. / Biochimie 91 (2009) 596–603
Fig. 7. Schematized comparison of the active sites of H. sapiens mt-AspRS and E. coli AspRS in interaction with aspartyl-adenylate and its two analogs, Asp-AMS and aspartol-AMP.
The figure shows the chemical structure of aspartyl-adenylate and the structural changes in Asp-AMS (in red) and aspartol-AMP (in magenta). The dashed line is the computedproximity contour of the adenylates in the binding cavity of AspRSs. The diameter of the shadowed blue circles is proportional to exposure of adenylate atoms to the solvent. Theamino acids forming the catalytic cavity are shown circled in three-letter abbreviations and numbering as in E. coli and human mitochondrial AspRSs (colored black and blue,respectively). The 10 amino acids in E. coli AspRS that play a role in catalysis , and present in human mt-AspRS, are displayed on a green background. Notice that Lys198 andArg489 were also predicted by free energy simulations to be the main contributors for the specificity of aspartate recognition by AspRSs . Conserved residues in all AspRSs are inbold, the other amino acids being semi-conserved and characteristic of bacterial- and mt-AspRSs. The green and blue arrows show respectively the amino acid side chains orbackbones predicted by the docking simulations to hydrogen bond with atoms from the adenylates. Note that several putative bonds are not possible with the analogs. (Forinterpretation of color in this figure, the reader is referred to the web version of this article.)
be kept in mind that the aminoacylation reaction is a two-step
a stronger effect on bacterial AspRSs (E. coli and P. aeruginosa) than
process including (i) formation of the aminoacyl-adenylate and (ii)
on human cytosolic AspRS. Here, for the first time, a very strong
transfer of the amino acid to the tRNA. Accordingly, the Ki value is
inhibition by Asp-AMS of a human mitochondrial synthetase has
not necessarily equal to the dissociation constant of the inhibitor
been measured. These data suggest that medical applications of
for the enzyme/inhibitor complex. Different relative rates of the
aaRS substrate analogs as inhibitors of pathogens could potentially
two steps of the reaction may account at least in part for the
affect the host mitochondrial enzymes. Since mitochondria are the
difference between the Ki values observed with mt-AspRS and cyt-
powerhouse of eukaryal cells, side effects can indeed not be ruled
AspRS. In regard to further structural investigations, we noticed
out. However, the toxicity of adenylate analogs in vivo is difficult to
that the strong binding of Asp-AMS, that differentiates the mt-
predict precisely since it will depend also on the ability of the drug
enzyme from other AspRSs, decreases its propensity to aggregate
to cross the mitochondrial membranes and further on the intra
and increases its solubility (not shown). Such a property, also found
mitochondrial concentration of free amino acid competing with the
in the case of human mt-TyrRS , becomes a positive hint
drug for the active site of the synthetase. Additional investigations
towards successful crystallization assays.
need to be performed to understand the contribution of theseparameters in detail.
The dramatic adaptation of pathogens to antibiotics calls for
new target macromolecules and new types of inhibitors. Along
M. Frugier is acknowledged for the generous gift of human cyt-
evolution, aaRSs acquired subtle differences in their active site,
AspRS. This work was supported by a collaborative France-Que´bec
making this family of macromolecules attractive targets in such
grant (project #61.103). We thank also Centre National de la
a strategy . Efficient inhibition by adenylate analogs has
Recherche Scientifique (CNRS) including a PICS project, Universite´
already been obtained for bacterial AspRS IleRS
Louis Pasteur Strasbourg (ULP), Association Française contre les
MetRS , GluRS and GlnRS Our present data confirm
Myopathies (AFM) in France and the Fonds Que´be´cois de Recherche
and extend a differential sensitivity of AspRSs from various
sur la Nature et les Technologies (FQRNT) in Canada for financial
organisms to aspartyl-adenylate analogs. Asp-AMS is the most
support. M.M. received a fellowship from the French Ministe re de
active inhibitor with Ki values in the nanomolar range, with
l'Enseignement Supe´rieur et de la Recherche.
M. Messmer et al. / Biochimie 91 (2009) 596–603
K. Arnold, L. Bordoli, J. Kopp, T. Schwede, The SWISS-MODEL workspace:a web-based environment for protein structure homology modelling, Bio-informatics 22 (2006) 195–201.
I.E. Scheffler, Mitochondria, second ed. Willey-Liss, 2007.
R.J. Rosenfeld, D.S. Goodsell, R.A. Musah, G.M. Morris, D.B. Goodin, A.J. Olson,
L. Bonnefond, A. Fender, J. Rudinger-Thirion, R. Giege´, C. Florentz, M. Sissler,
Automated docking of ligands to an artificial active site: augmenting crys-
Towards the full set of human mitochondrial aminoacyl-tRNA synthetases:
tallographic analysis with computer modeling, J. Comput. Aided Mol. Des. 17
characterization of AspRS and TyrRS, Biochemistry 44 (2005) 4805–4816.
(2003) 525–536.
M. Sissler, B. Lorber, M. Messmer, A. Schaller, J. Pu¨tz, C. Florentz, Handling
P. Berg, E.J. Offengand, An enzymatic mechanism for linking amino acids to
mammalian mitochondrial tRNAs and aminoacyl-tRNA synthetases for
RNA, Proc. Nat. Acad. Sci. U.S.A. 44 (1958) 78–86.
functional and structural characterization, Methods 44 (2008) 176–189.
J. Lapointe, R. Giege´, Transfer RNAs and aminoacyl-tRNA synthetases, in:
K. Shiba, P. Schimmel, H. Motegi, T. Noda, Human glycyl-tRNA synthetase.
H. Trachsel (Ed.), Translation in Eukaryotes, CRC Presss Inc, Boca Raton, FL,
Wide divergence of primary structure from bacterial counterpart and
1991, pp. 35–69.
species-specific aminoacylation, J. Biol. Chem. 269 (1994) 30049–30055.
R. Giege´, The early history of tRNA recognition by aminoacyl-tRNA synthe-
S.J. Mudge, J.H. Williams, H.J. Eyre, G.R. Sutherland, P.J. Cowan, D.A. Power,
tases, J. Biosci. 31 (2006) 477–488.
Complex organisation of the 50-end of the human glycine tRNA synthetase
M. Ibba, D. So¨ll, Aminoacyl-tRNA synthesis, Annu. Rev. Biochem. 69 (2000)
gene, Gene 209 (1998) 45–50.
E. Tolkunova, H. Park, J. Xia, M.P. King, E. Davidson, The human lysyl-tRNA
M. Ibba, C. Francklyn, S. Cusack (Eds.), The Aminoacyl-tRNA Synthetases,
synthetase gene encodes both the cytoplasmic and mitochondrial enzymes
Landes Bioscience, 2005.
by means of an unusual splicing of the primary transcript, J. Biol. Chem. 275
K.B. Jacobson, Reaction of aminoacyl-tRNA synthetases with heterologous
tRNA's, Prog. Nucl. Acid Res. Mol. Biol. 11 (1971) 461–488.
L. Margulis, Origin of Eukaryotic Cells, New Haven Yale University Press, 1970.
A.R. Fersht, J.P. Shi, J. Knill-Jones, D.M. Lowe, A.J. Wilkinson, D.M. Blow,
M.W. Gray, G. Burger, B.F. Lang, The origin and early evolution of mito-
P. Brick, P. Carter, M.M. Waye, G. Winter, Hydrogen bonding and biological
chondria, Genome Biol. 2 (2001) 1018.1–1018.5.
specificity analysed by protein engineering, Nature 314 (1985) 235–238.
B. Brindefalk, J. Viklund, D. Larsson, M. Thollesson, S.G. Andersson, Origin and
J. Pu¨tz, J.D. Puglisi, C. Florentz, R. Giege´, Additive, cooperative and anti-
evolution of the mitochondrial aminoacyl-tRNA synthetases, Mol. Biol. Evol.
cooperative effects between identity nucleotides of a tRNA, EMBO J. 12
24 (2007) 743–756.
(1993) 2949–2957.
L. Bonnefond, M. Frugier, E. Touze´, B. Lorber, C. Florentz, R. Giege´, C. Sauter,
B. Lorber, D. Kern, R. Giege´, J.-P. Ebel, Covalent attachment of aspartic acid to
J. Rudinger-Thirion, Crystal structure of human mitochondrial tyrosyl-tRNA
yeast aspartyl-tRNA synthetase, FEBS Lett. 146 (1982) 59–64.
synthetase reveals common and idiosyncratic features, Structure 15 (2007)
D. Kern, B. Lorber, Y. Boulanger, R. Giege´, A peculiar property of aspartyl-
tRNA synthetase from bakers yeast: chemical modification of the protein by
A. Fender, C. Sauter, M. Messmer, J. Pu¨tz, R. Giege´, C. Florentz, M. Sissler, Loss
the enzymatically synthesized aminoacyl adenylate, Biochemistry 24 (1985)
of a primordial identity element for a mammalian mitochondrial amino-
acylation system, J. Biol. Chem. 281 (2006) 15980–15986.
D. Bernard, P.M. Akochy, S. Bernier, O. Fisette, O.C. Brousseau, R. Cheˆnevert,
L. Bonnefond, M. Frugier, R. Giege´, J. Rudinger-Thirion, Human mitochondrial
P.H. Roy, J. Lapointe, Inhibition by
TyrRS disobeys the tyrosine identity rules, RNA 11 (2005) 558–562.
L-aspartol adenylate of a nondiscriminating
aspartyl-tRNA synthetase reveals differences between the interactions of its
Y. Kumazawa, H. Himeno, K.-I. Miura, K. Watanabe, Unilateral aminoacylation
active site with tRNA(Asp) and tRNA(Asn), J. Enzyme Inhib. Med. Chem. 22
specificity between bovine mitochondria and eubacteria, J. Biochem. 109
(2007) 77–82.
(1991) 421–427.
S. Cusack, C. Berthet-Colominas, M. Hartlein, N. Nassar, R. Leberman, A
J. Tao, P. Schimmel, Inhibitors of aminoacyl-tRNA synthetases as novel anti-
second class of synthetase structure revealed by X-ray analysis of Escherichia
infectives, Exp. Opin. Invest. Drugs 9 (2000) 1767–1775.
coli seryl-tRNA synthetase at 2.5 Å, Nature 347 (1990) 249–255.
R. Cheˆnevert, S. Bernier, J. Lapointe, Inhibitors of aminoacyl-tRNA synthetases
G. Eriani, M. Delarue, O. Poch, J. Gangloff, D. Moras, Partition of tRNA
as antibiotics and tools for structural and mechanistic studies, in: J. Lapointe,
synthetases into two classes based on mutually exclusive sets of sequence
L. Brakier-Gringas (Eds.), Translation Mechanisms, Landes Bioscience, 2003,
motifs, Nature 347 (1990) 203–206.
pp. 416–428.
R. Giege´, B. Rees, Aspartyl-tRNA synthetases, in: M. Ibba, C. Francklyn,
S. Kim, C. Lee, E.C. Choi, S.Y. Choi, Aminoacyl-tRNA synthetases and their
S. Cusack (Eds.), Aminoacyl-tRNA synthetases, Landes Biosciences, 2005, pp.
inhibitors as a novel family of antibiotics, Appl. Microbiol. Biotechnol. 61
(2003) 278–288.
E. Schmitt, L. Moulinier, S. Fujiwara, T. Imanaka, J.-C. Thierry, D. Moras,
J. Pohlmann, H. Brotz-Oesterhelt, New aminoacyl-tRNA synthetase inhibitors
Crystal structure of aspartyl-tRNA synthetase from Pyrococcus kodakaraensis
as antibacterial agents, Curr. Drug Targets Infect. Disord. 4 (2004) 261–272.
KOD: archeon specificity and catalytic mechanism of adenylate formation,
M. Torchala, M. Hoffmann, IA, Database of known ligands of aminoacyl-tRNA
EMBO J. 17 (1998) 5227–5237.
synthetases, J. Comput. Aided Mol. Des. 21 (2007) 523–525.
L. Ador, A. Camasses, P. Erbs, J. Cavarelli, D. Moras, J. Gangloff, G. Eriani,
A. Fuller, Pseudomonic acid: an antibiotic produced by Pseudomonas
Active site mapping of yeast aspartyl-tRNA synthetase by in vivo selection
fluorescens, Nature 234 (1971) 416–417.
of enzyme mutations lethal for cell growth, J. Mol. Biol. 288 (1999)
J. Hughes, G. Mellows, Inhibition of isoleucyl-transfer ribonucleic acid synthe-
tase in Escherichia coli by pseudomonic acid, Biochem. J. 176 (1978) 305–318.
R. Giege´, Toward a more complete view of tRNA biology, Nat. Struct. Mol.
R.O. Nicholas, V. Berry, P.A. Hunter, J.A. Kelly, The antifungal activity of
Biol. 15 (2008) 1007–1014.
mupirocin, J. Antimicrob. Chemother. 43 (1999) 579–582.
M. Sissler, J. Pu¨tz, F. Fasiolo, C. Florentz, Mitochondrial aminoacyl-tRNA
S. Bernier, P.M. Akochy, J. Lapointe, R. Cheˆnevert, Synthesis and aminoacyl-
synthetases, in: M. Ibba, C. Francklyn, S. Cusack (Eds.), Aminoacyl-tRNA
tRNA synthetase inhibitory activity of aspartyl adenylate analogs, Bioorg.
synthetases, Landes Biosciences, 2005, pp. 271–284.
Med. Chem. 13 (2005) 69–75.
A.J. Pope, M. McVey, K. Fantom, K.J. Moore, Effects of substrate and inhibitor
J. Lapointe, S. Levasseur, D. Kern, Glutamyl-tRNA synthetase from Escherichia
binding on proteolysis of isoleucyl-tRNA synthetase from Staphylococcus
coli, Methods Enzymol. 113 (1985) 42–49.
aureus, J. Biol. Chem. 273 (1998) 31702–31706.
C. Balg, S.P. Blais, S. Bernier, J.L. Huot, M. Couture, J. Lapointe, R. Cheˆnevert,
C.F. Crasto, A.K. Forrest, T. Karoli, D.R. March, L. Mensah, P.J. O'Hanlon,
Synthesis of beta-ketophosphonate analogs of glutamyl and glutaminyl
M.R. Nairn, M.D. Oldham, W. Yue, M.G. Banwell, C.J. Easton, Synthesis and
adenylate, and selective inhibition of the corresponding bacterial aminoacyl-
activity of analogues of the isoleucyl tRNA synthetase inhibitor SB-203207,
tRNA synthetases, Bioorg. Med. Chem. 15 (2007) 295–304.
Bioorg. Med. Chem. 11 (2003) 2687–2694.
M. Ruff, S. Krishnaswamy, M. Boeglin, A. Poterszman, A. Mitschler,
J. Lee, S.E. Kim, J.Y. Lee, S.Y. Kim, S.U. Kang, S.H. Seo, M.W. Chun, T. Kang,
A. Podjarny, B. Rees, J.-C. Thierry, D. Moras, Class II aminoacyl transfer RNA
S.Y. Choi, H.O. Kim, N-Alkoxysulfamide, N-hydroxysulfamide, and sulfamate
synthetases: crystal structure of yeast aspartyl-tRNA synthetase complexed
analogues of methionyl and isoleucyl adenylates as inhibitors of methionyl-
with tRNAAsp, Science 252 (1991) 1682–1689.
tRNA and isoleucyl-tRNA synthetases, Bioorg. Med. Chem. Lett. 13 (2003)
S. Eiler, A.C. Dock-Bregeon, L. Moulinier, J.-C. Thierry, D. Moras, Synthesis of
aspartyl-tRNAAsp in Escherichia coli – a snapshot of the second step, EMBO J.
G. Archontis, T. Simonson, D. Moras, M. Karplus, Specific amino acid recog-
18 (1999) 6532–6541.
nition by aspartyl-tRNA synthetase studied by free energy simulations, J. Mol.
A. Sali, T.L. Blundell, Comparative protein modelling by satisfaction of spatial
Biol. 275 (1998) 823–846.
restraints, J. Mol. Biol. 234 (1993) 779–815.
Source: http://cj.sauter.free.fr/xtal/pdf/messmer-biochimie09.pdf
© SYMPHONYA Emerging Issues in Management, n. 2, 2014 symphonya.unimib.it Preventing Corruption in Africa: Emerging Challenges in the Mining Sector of the Democratic Republic of Congo* Cosetta Pepe**, Jean Marie Mushagalusa Nshombo***, Mario Risso**** The globalisation of economies and markets, brings out the full importance of the
6th Annual North Park University Undergraduate Research Symposium Tuesday, April 17, 2012 North Park University Chicago, Illinois Dr. Rachel Schmale Session 1 John-Tyler Carlson Session 2 Closing Remarks 5:20–5:25 pm Dr. Matthew Schau Following the symposium: Discussion and dinner (served at 5:45 pm) for presenters and advisors in Olssson Lounge, Seminary Building.






